Give a Specific Example of How the Introduction of a Gene Into a Bacterium Can Change the Phenotype
GloFish are the get-go transgenic animals available to the American public. But what'due south the biotechnology behind them?

Figure 1: The multicolored GloFish®.
Courtesy of world wide web.glofish.com. All rights reserved. ![]()
"Seeing is believing with GloFish. They are admittedly stunning!" The preceding is some of the marketing material you'd read if y'all visited the GloFish website (GloFish, 2008). Beauty may be in the eye of the beholder, just nearly everyone would agree that these beginning—and, so far, only—transgenic animals made available to the full general public in the U.s.a. (except in California, awaiting a formal review of their potential consequence on the environs) are a worthy conversation piece. A transgenic, or genetically modified, organism is one that has been altered through recombinant Deoxyribonucleic acid technology, which involves either the combining of DNA from different genomes or the insertion of foreign DNA into a genome. GloFish (Figure i) are a type of transgenic zebrafish ( Danio rerio ) that take been modified through the insertion of a light-green fluorescent protein (gfp) cistron. Non all GloFish are green, however. Rather, there are several gfp cistron constructs, each encoding a different colored phenotype, from fluorescent xanthous to fluorescent ruddy.
Currently, GloFish are the merely recombinant-DNA animal that has been approved for homo "apply" by the U.S. Nutrient and Drug Assistants. Their approval has raised of import questions virtually whether, and to what extent, genetically modified animals should be made bachelor to consumers. But how were scientists able to create these engineered organisms in the start place? Similar and then many genetic technologies used today, recombinant DNA technology had its origins in the late 1960s and early 1970s. Past the 1960s, scientists had already learned that cells repair Dna breaks by reuniting, or recombining, the broken pieces. Thus, it was only a matter of fourth dimension before researchers identified the raw biological ingredients necessary for recombination, figured out how those ingredients functioned together, and then tried to govern the recombining process themselves.
Early on Experiments Provide the Basis for Recombinant Organisms
Although recombinant Deoxyribonucleic acid applied science first emerged in the 1960s and 1970s, the basic principle of recombination had been discovered many years earlier. Indeed, in 1928, Frederick Griffith, an English medical officeholder studying the bacteria responsible for a pneumonia epidemic in London, first demonstrated what he termed "genetic transformation"; hither, living cells took upwardly genetic fabric released by other cells and became phenotypically "transformed" by the new genetic information. More than a decade later, Oswald Avery repeated Griffith's piece of work and isolated the transforming molecule, which turned out to be Deoxyribonucleic acid. These experiments showed that Deoxyribonucleic acid can be transferred from one prison cell to some other in the laboratory, thus irresolute the actual genetic phenotype of an organism.
Prior to these classic experiments, the idea that the genetic material was a specific chemical that could be modified and transferred into cells was certainly controversial. But before the explosion in recombinant DNA could begin, scientists would have to learn not only how to transfer Dna, but besides how to isolate and modify individual genes.
Fundamental Developments in Recombinant Deoxyribonucleic acid Applied science
Following these early experiments, four cardinal developments helped lead to construction of the first recombinant Dna organism (Kiermer, 2007). The first two developments revolved around how scientists learned to cutting and paste pieces of DNA from dissimilar genomes using enzymes. The latter 2 events involved the evolution of techniques used to transfer foreign DNA into new host cells.
Discovering the Cut-and-Paste Enzymes
The first major footstep forward in the power to chemically alter genes occurred when American biologist Martin Gellert and his colleagues from the National Institutes of Wellness purified and characterized an enzyme in Escherichia coli responsible for the bodily joining, or recombining, of separate pieces of Dna (Zimmerman et al., 1967). They chosen their observe "Deoxyribonucleic acid-joining enzyme," and this enzyme is now known as Deoxyribonucleic acid ligase. All living cells use some version of Dna ligase to "glue together" short strands of DNA during replication. Using E. coli excerpt, the researchers adjacent showed that only in the presence of ligase was it possible to repair unmarried-stranded breaks in λ phage DNA. (Discovered in 1950 by American microbiologist Esther Lederberg, λ phage is a virus particle that infects Eastward. coli.) More specifically, they showed that the enzyme was able to form a 3'-5'-phosphodiester bail between the 5'-phosphate end of the last nucleotide on 1 Deoxyribonucleic acid fragment and the iii'-OH finish of the concluding nucleotide on an adjacent fragment. The identification of Deoxyribonucleic acid ligase was the kickoff of several key steps that would eventually empower scientists to attempt their own recombination experiments—experiments that involved non just recombining the DNA of a unmarried individual, just recombining Deoxyribonucleic acid from dissimilar individuals, including different species.
A 2d major footstep forward in factor modification was the discovery of restriction enzymes, which cleave Deoxyribonucleic acid at specific sequences. These enzymes were discovered at approximately the same time as the first DNA ligases by Swiss biologist Werner Arber and his colleagues while they were investigating a phenomenon called host-controlled restriction of bacteriophages. Bacteriophages are viruses that invade and frequently destroy their bacterial host cells; host-controlled restriction refers to the defense mechanisms that bacterial cells have evolved to deal with these invading viruses. Arber'south team discovered that i such mechanism is enzymatic action provided by the host prison cell. The squad named the responsible enzymes "restriction enzymes" because of the way they restrict the growth of bacteriophages. These scientists were besides the first to demonstrate that restriction enzymes damage invading bacteriophages past cleaving the phage Dna at very specific nucleotide sequences (now known as restriction sites). The identification and characterization of restriction enzymes gave biologists the means to cut specific pieces of DNA required (or desired) for subsequent recombination.
Inserting Foreign DNA into a New Host Cell
Although Griffith and Avery had had demonstrated the ability to transfer foreign genetic textile into cells decades before, this "transformation" was very inefficient, and it involved "natural" rather than manipulated Dna. Simply in the 1970s did scientists begin to use vectors to efficiently transfer genes into bacterial cells. The kickoff such vectors were plasmids, or small DNA molecules that live naturally inside bacterial cells and replicate separately from a bacterium's chromosomal DNA.
Plasmids' utility as a Deoxyribonucleic acid shuttle, or vector, was discovered by Stanford University biochemist Stanley Cohen. Scientists had already established that some bacteria had what were known as antibiotic resistance factors, or R factor-plasmids that replicated independently inside the bacterial cell. Only scientists knew little about how the different R factor genes functioned. Cohen thought that if there were an experimental organization for transforming host bacterial cells with these R-gene DNA molecules, he and other researchers might exist able to improve sympathise R-factor biology and figure out exactly what it was almost these plasmids that made bacteria antibiotic-resistant. He and his colleagues developed that system by demonstrating that calcium chloride-treated Due east. coli tin be genetically transformed into antibiotic-resistant cells by the improver of purified plasmid DNA (in this case, purified R-factor DNA) to the bacteria during transformation (Cohen et al., 1972).
Recombinant Plasmids in Leaner
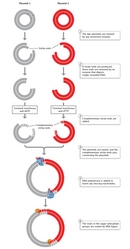
The post-obit year, Stanley Cohen and his colleagues were also the first to construct a novel plasmid DNA from two separate plasmid species which, when introduced into E. coli, possessed all the nucleotide base of operations sequences and functions of both parent plasmids. Cohen's team used brake endonuclease enzymes to cleave the double-stranded Deoxyribonucleic acid molecules of the two parent plasmids. The team next used Dna ligase to rejoin, or recombine, the DNA fragments from the two different plasmids (Figure ii). Finally, they introduced the newly recombined plasmid Deoxyribonucleic acid into East. coli. The researchers were able to bring together two Dna fragments from completely different plasmids because, as they explained, "the nucleotide sequences broken are unique and self-complementary so that Deoxyribonucleic acid fragments produced by 1 of these enzymes tin associate by hydrogen-bonding with other fragments produced past the aforementioned enzyme" (Cohen et al., 1973).

The same could be said of whatever DNA—non merely plasmids—from two unlike species. This universality—the capacity to mix and match Dna from different species, because Deoxyribonucleic acid has the same structure and role in all species and considering restriction and ligase enzymes cut and paste the same means in different genomes—makes recombinant Deoxyribonucleic acid biology possible.
Today, the E. coli λ bacteriophage is 1 of the nigh widely used vectors used to deport recombinant Dna into bacterial cells. This virus makes an excellent vector because about ane-tertiary of its genome is considered nonessential, meaning that it can be removed and replaced by foreign Dna (i.e., the Deoxyribonucleic acid being inserted). As illustrated in Figure iii, the nonessential genes are removed past restriction enzymes (the specific restriction enzyme EcoRI is shown in the figure), the foreign DNA inserted in their place, and then the concluding recombinant Dna molecule is packaged into the virus'south protein coat and prepped for introduction into its host cell.
Vectors Used in Mammalian Cells
A fourth major pace forward in the field of recombinant DNA engineering was the discovery of a vector for efficiently introducing genes into mammalian cells. Specifically, researchers learned that recombinant Dna could be introduced into the SV40 virus, a pathogen that infects both monkeys and humans. Indeed, in 1972, Stanford Academy researcher Paul Berg and his colleagues integrated segments of λ phage Dna, likewise as a segment of E. coli Deoxyribonucleic acid containing the galactose operon, into the SV40 genome. (The E. coli galactose operon is a cluster of genes that plays a role in galactose sugar metabolism.) The significance of their achievement was its sit-in that recombinant DNA technologies could be practical to essentially any Dna sequences, no matter how distantly related their species of origin. In their words, these researchers "developed biochemical techniques that are more often than not applicable for joining covalently whatever two DNA molecules" (Jackson et al., 1972). While the scientists didn't really introduce foreign Deoxyribonucleic acid into a mammalian jail cell in this experiment, they provided (proved) the means to do so.
Recombinant Deoxyribonucleic acid Applied science Creates Recombinant Animals
The showtime bodily recombinant animal cells weren't developed until nearly a decade afterward the inquiry conducted past Berg's squad, and most of the early studies involved mouse cells. In 1981, for instance, Franklin Costantini and Elizabeth Lacy of the University of Oxford introduced rabbit DNA fragments containing the developed beta globin factor into murine (mouse) germ-line cells (Costantini & Lacy, 1981). (The beta globins are a family of polypeptides that serve every bit the subunits of hemoglobin molecules.) Another group of scientists had demonstrated that foreign genes could be successfully integrated into murine somatic cells, merely this was the first demonstration of their integration into germ cells. In other words, Costantini and Lacy were the starting time to engineer an unabridged recombinant animal (albeit with relatively low efficiency).
Interestingly, non long after the publication of his squad'south 1972 study, Paul Berg led a voluntary moratorium in the scientific community against sure types of recombinant Deoxyribonucleic acid research. Clearly, scientists take always been aware that the ability to manipulate the genome and mix and match genes from unlike organisms, even different species, raises firsthand and serious questions about the potential hazards and risks of doing so—implications nonetheless being debated today.
Since these early studies, scientists have used recombinant DNA technologies to create many different types of recombinant animals, both for scientific study and for the profitable manufacturing of man proteins. For instance, mice, goats, and cows take all been engineered to create medically valuable proteins in their milk; moreover, hormones that were once isolated only in pocket-size amounts from human cadavers tin now be mass-produced by genetically engineered cells. In fact, the entire biotechnology industry is based upon the ability to add new genes to cells, plants, and animals Equally scientists discover important new proteins and genes, these technologies will continue to course the foundation of future generations of discoveries and medical advances.
References and Recommended Reading
Cohen, Due south. N., et al. Nonchromosomal antibiotic resistance in bacteria: Genetic transformation of Escherichia coli by R-cistron DNA. Proceedings of the National Academy of Sciences 69, 2110–2114 (1972)
———. Construction of biologically functional bacterial plasmids in vitro. Proceedings of the National Academy of Sciences 70, 3240–3244 (1973)
Costantini, F., & Lacy, E. Introduction of a rabbit beta-globin gene into the mouse germ line. Nature 294, 92–94 (1981) (link to article)
Crea, R., et al. Chemical synthesis of genes for human insulin. Proceedings of the National University of Sciences 75, 5765–5769 (1978)
GloFish. GloFish abode folio. www.glofish.com (Accessed July 3, 2008)
Jackson, D. A., et al. Biochemical method for inserting new genetic data into Deoxyribonucleic acid of simian virus 40: Circular SV40 Deoxyribonucleic acid molecules containing lambda phage genes and the galactose operon of Escherichia coli. Proceedings of the National Academy of Sciences 69, 2904–2909 (1972)
Kiermer, V. The dawn of recombinant Deoxyribonucleic acid. Nature Milestones: Dna Technologies, http://www.nature.com/milestones/miledna/total/miledna02.html (2007) (link to article)
Miller, H. I. FDA on transgenic animals—A dog's breakfast? Nature Biotechnology 26, 159–160 (2008) (link to article)
Zimmerman, S. B., et al. Enzymatic joining of Dna strands: A novel reaction of diphosphopyridine nucleotide. Proceedings of the National University of Sciences 57, 1841–1848 (1967)
Source: http://www.nature.com/scitable/topicpage/recombinant-dna-technology-and-transgenic-animals-34513
0 Response to "Give a Specific Example of How the Introduction of a Gene Into a Bacterium Can Change the Phenotype"
Postar um comentário